Performance Engineering in Practice
Algorithm Engineering — T01
Performance Engineering in Practice
We know why code gets slow. This tutorial is about how to react.
Block 1: Find the bottleneck. Block 2: Fix the bottleneck.
Block 1: Find the Bottleneck
Before tuning anything, we need two guardrails: Amdahl’s law and “premature optimization is the root of all evil.”
Amdahl’s law: know the ceiling
If fraction \(p\) of runtime can be improved by factor \(s\), then
\[ \text{speedup} = \frac{1}{(1-p) + \frac{p}{s}} \]
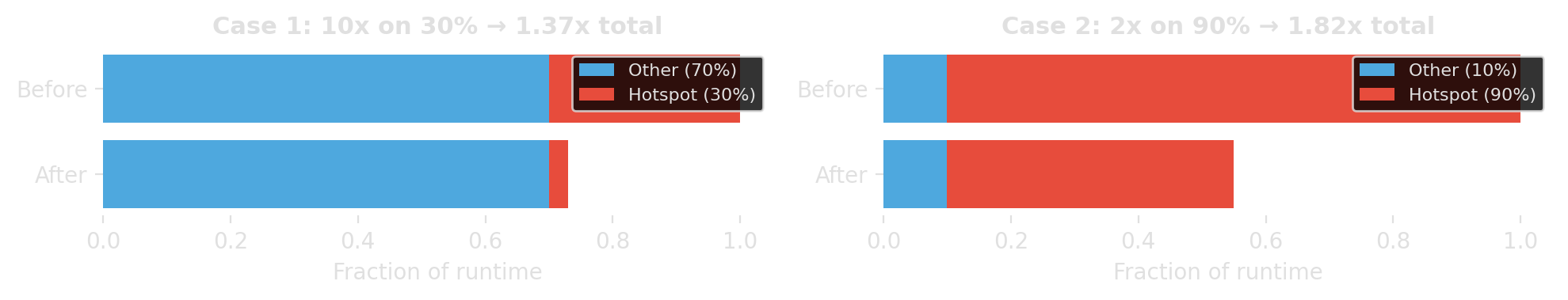
Speed up the part that dominates runtime, not the part that looks embarrassing.
“Premature optimization is the root of all evil”
“We should forget about small efficiencies, say about 97% of the time: premature optimization is the root of all evil. Yet we should not pass up our opportunities in that critical 3%.”
Donald Knuth, “Structured Programming with go to Statements”, ACM Computing Surveys, 1974. (Knuth 1974)
This quote does not mean “performance does not matter.” It means: optimize after you know where the real bottleneck is.

Photo: Alex Handy, CC BY-SA 2.0
Observability: Basic Logging
Before profilers, try the simplest thing that could work.
Start with phase markers
import logging
logging.basicConfig(
level=logging.INFO,
format="%(asctime)s %(message)s",
datefmt="%H:%M:%S",
)
log = logging.getLogger("solver")
log.info("Loading instance: nodes=%d, edges=%d", n_nodes, n_edges)
# ... loading code ...
log.info("Computing greedy upper bound")
# ... heuristic code ...
log.info("Starting solver")
# ... solver code ...
log.info("Done")The logger already adds timestamps. That is often enough to identify the slow phase.
What the timestamps reveal
16:58:12 Loading instance: nodes=500, edges=12340
16:58:12 Building graph
16:58:12 Graph built: nodes=500, edges=12340, density=0.10
16:58:12 Computing greedy upper bound
16:58:16 Upper bound found: colors=12
16:58:16 Starting solver
16:58:18 Solver finished: colors=8, optimal=True
16:58:18 Writing solution
16:58:18 DoneThe “quick” heuristic is the bottleneck — 4 seconds, while the actual solver took only 2. That is where we need to speed things up.
Add structural context
Without input-size context, runtime logs are much harder to interpret.
Log structural statistics at natural boundaries: input size, graph size, iteration counts, constraint counts.
Controlling verbosity per component
# Each module gets its own logger
log_model = logging.getLogger("solver.model")
log_heur = logging.getLogger("solver.heuristic")
# Use levels to express importance
log_model.info("Variables: %d, constraints: %d", n_vars, n_cons)
log_model.debug("Constraint matrix density: %.4f", density)
log_heur.warning("Greedy coloring exceeded timeout")
# Dial verbosity up or down per component
log_model.setLevel(logging.DEBUG) # loud — investigating model size
log_heur.setLevel(logging.INFO) # normalLeave log calls in the code permanently. Dial verbosity up when investigating, down for production runs.
Guard expensive log computations
log.debug(...) skips formatting, but Python still evaluates all arguments before the call.
Guard with isEnabledFor() when an argument is expensive to compute.
Good code has logging — especially in production
- phase markers let you pinpoint issues without reproducing them locally
- add progress logs inside long loops (every N items, not every item)
- use them to see scaling behavior and estimate remaining runtime
Logging tells you which phase is slow. The structural stats often tell you why at a high level. Only then do you reach for finer-grained tools.
Profiling
Logging tells us where to look. Profilers tell us how the cost is distributed inside the code.
Two big families
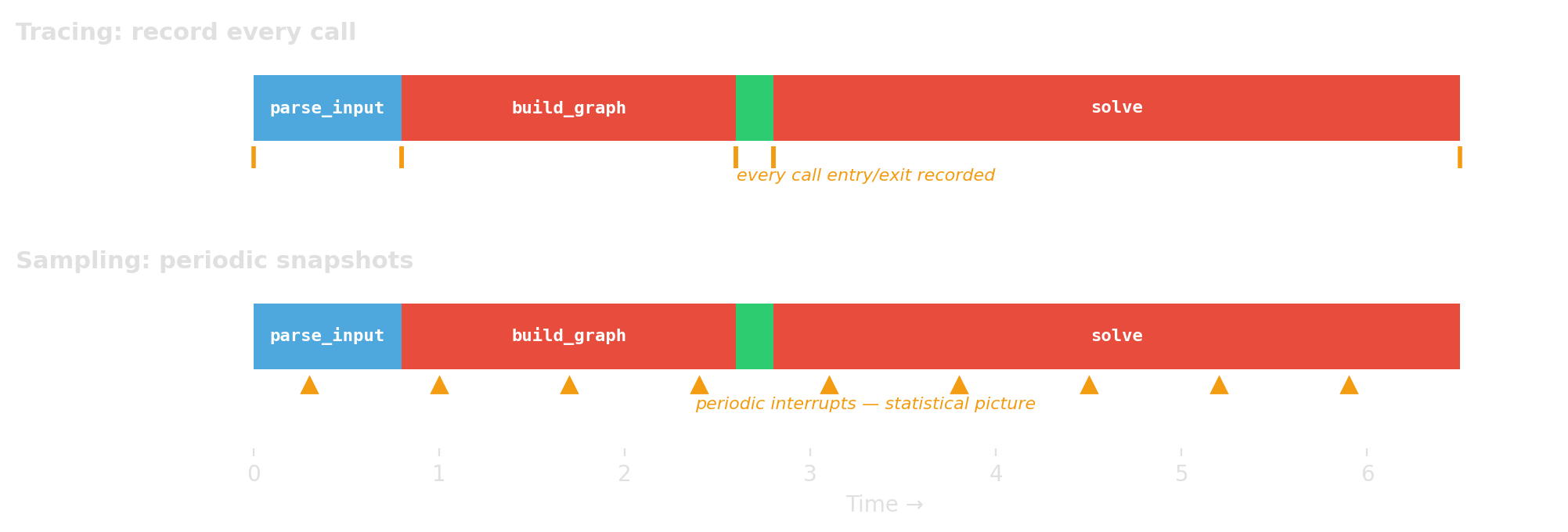
Tracing / instrumentation
- record every call or event
- exact call counts, higher overhead
- Python example:
cProfile
Sampling
- interrupt periodically
- statistical estimate, lower overhead
- Python example:
Scalene
What does “time” even mean?
- CPU time: time while the program is actually running on CPU
- wall-clock time: elapsed real time, including waiting
- on-CPU profiles can miss I/O waits and blocking
- mixed Python/native systems may need a split like Python vs native vs system
If a Python program spends a lot of time waiting or inside native libraries, the profiler must match that reality.
A practical rule of thumb
- start with logging to find the slow phase
- use Scalene for line-level Python vs native insight
- use cProfile when you want exact call counts and call paths
- if the hotspot is in native code, use native profilers (
perf, VTune)
Start broad and cheap. Go narrower only where the evidence points.
cProfile: when you want exact call counts
build_graph called 1225 times — if you expected once, that is the bug. Trust call counts from cProfile. Treat timing as approximate; tracing adds overhead.
cProfile: targeted profiling
# Profile just one function call
import cProfile
cProfile.run("build_graph(data)", sort="tottime")
# Profile a specific section with the context manager
with cProfile.Profile() as pr:
result = build_graph(data)
solution = solve(result)
pr.print_stats(sort="tottime") # print to stdout
pr.dump_stats("profile.dat") # save for later analysis 2374431 function calls (828274 primitive calls) in 0.757 seconds
Ordered by: internal time
ncalls tottime percall cumtime percall filename:lineno(function)
206534 0.514 0.000 0.514 0.000 cp_model.py:537(add_all_different)
1752679/206535 0.214 0.000 0.221 0.000 profile_solver.py:39(_extend)
206535 0.012 0.000 0.232 0.000 profile_solver.py:34(enumerate_all_cliques)
1 0.010 0.010 0.242 0.242 cp_model.py:333(new_int_var)Scalene: when you want line-level insight (wall-clock, Python/native split)
--program-path .— only profile your code, not libraries (crucial!)--html --output profile.html— generate a browsable HTML report- without
--program-path, the output is dominated by library internals
Reading Scalene output
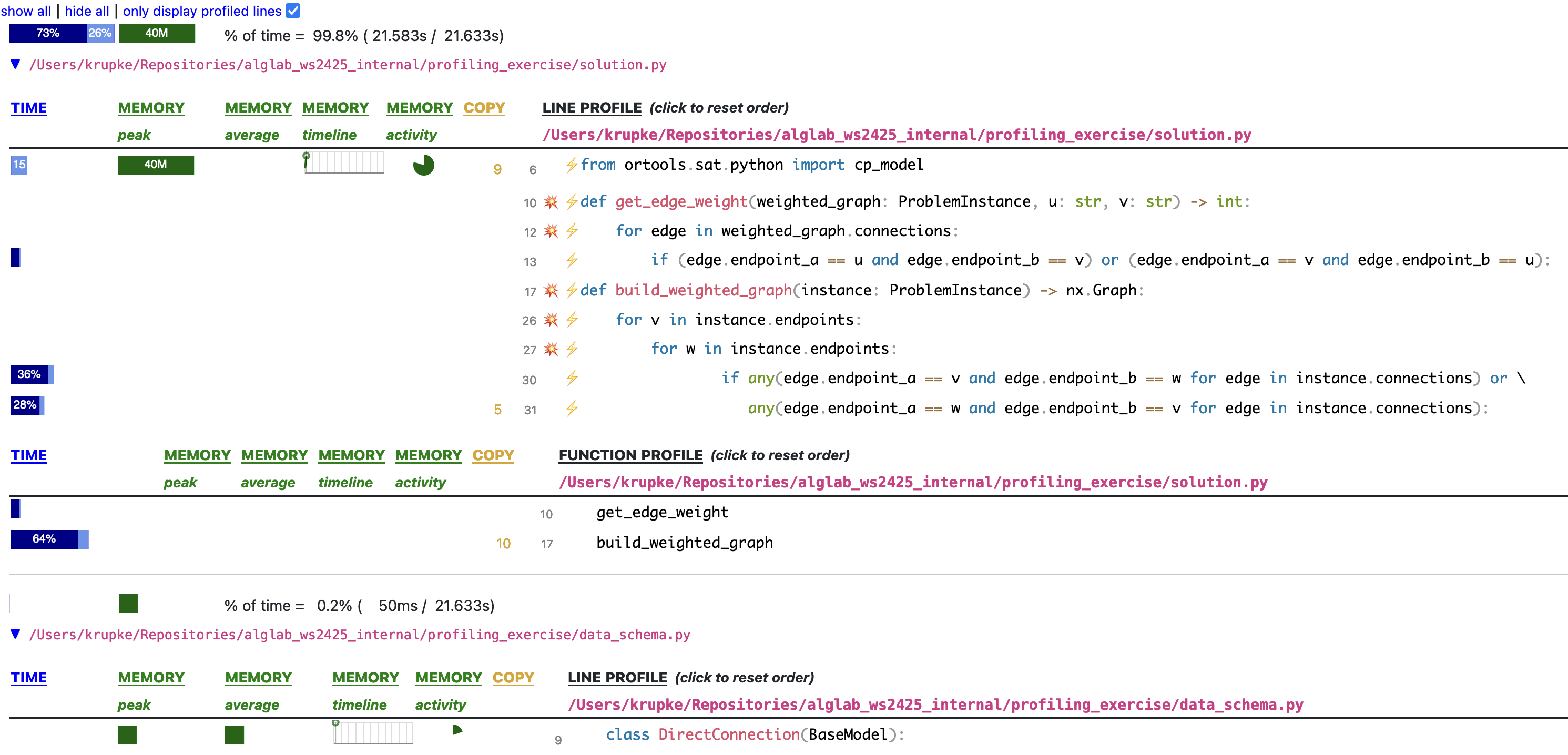

For each file, Scalene shows per-line and per-function time. Here, build_weighted_graph takes 64% — and line 30 alone takes 36%.
C++ profiling with perf
Profiling C++ is more painful than Python — but the payoff is a view all the way down to the hardware.
perfreads the CPU’s own counters: cycles, cache misses, branch mispredictions- You see not just where the CPU spent time, but why it spent it there
- No code changes, no recompile beyond
-gfor symbols
Walkthrough example: examples/perf_record/
perf answers two questions
- What kind of bottleneck is this? — compute, memory, branches, kernel, waiting
- count events across the run →
perf stat
- count events across the run →
- Where in the code does it land? — function, line, call path
- sample the program counter →
perf record
- sample the program counter →
Full program → perf record first to find the hotspot. Isolated benchmark → perf stat first to classify the bottleneck. But beware: an isolated benchmark may behave differently — warm caches, no memory pressure from surrounding code.
perf stat — classify
- IPC (instructions per cycle) is the first-order signal — 0.19 means the CPU is mostly stalled
- Read IPC together with total instructions and wall time — not alone
- Vectorizing can raise CPI while making code faster (fewer instructions, same work)
-r 5repeats the run and reports variance — use it for any measurement you’ll act on
Read the signature, not the number
Individual counters rarely diagnose. Patterns across counters do.
| Pattern | Counter signature | Likely cause |
|---|---|---|
| Compute-bound | High IPC, low miss rate | Instruction count / algorithm |
| Memory-bound | Low IPC, miss pressure high | Locality, data layout |
| Branch-heavy | Branch-miss % high on hot code | Unpredictable conditionals |
| Kernel-heavy | cycles:k comparable to cycles:u |
Syscalls, I/O path |
| Off-CPU | task-clock ≪ wall time |
Waiting, not computing |
Last row is the trap: if task-clock ≪ wall time, your program is blocked, not slow. You need off-CPU tools (perf record --off-cpu, perf sched, perf lock) — sampling harder won’t help.
perf record + perf report: find the hotspot
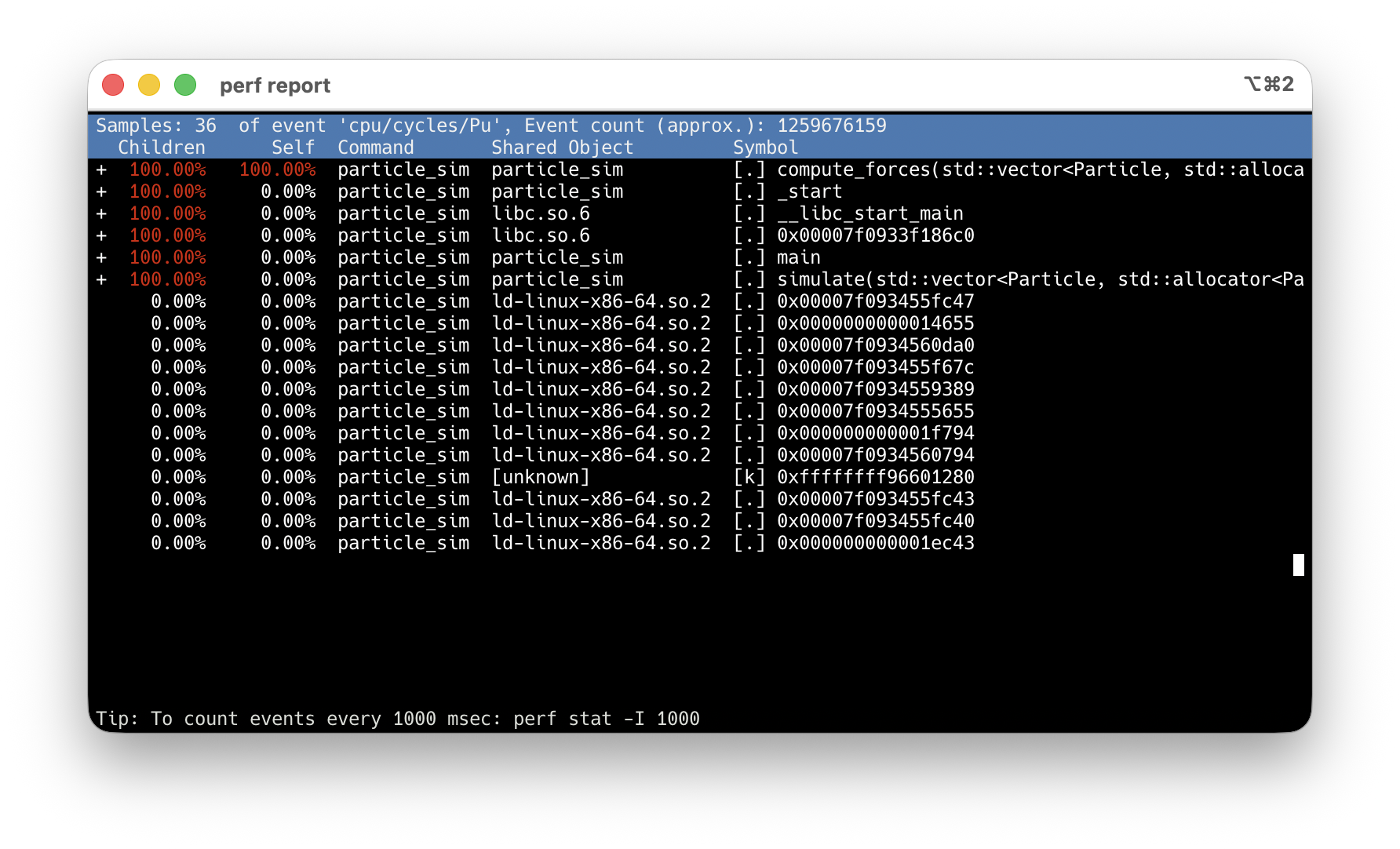
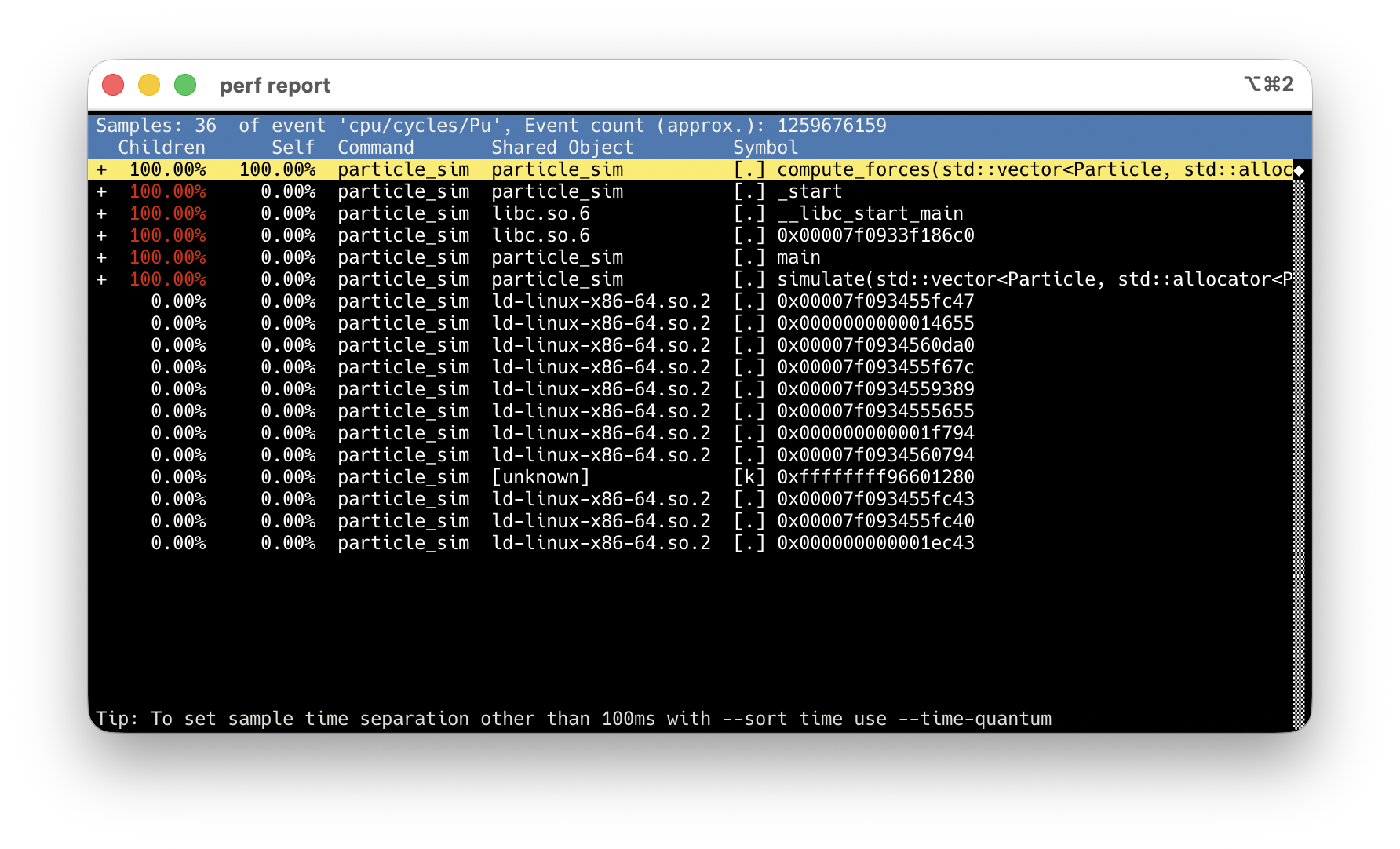
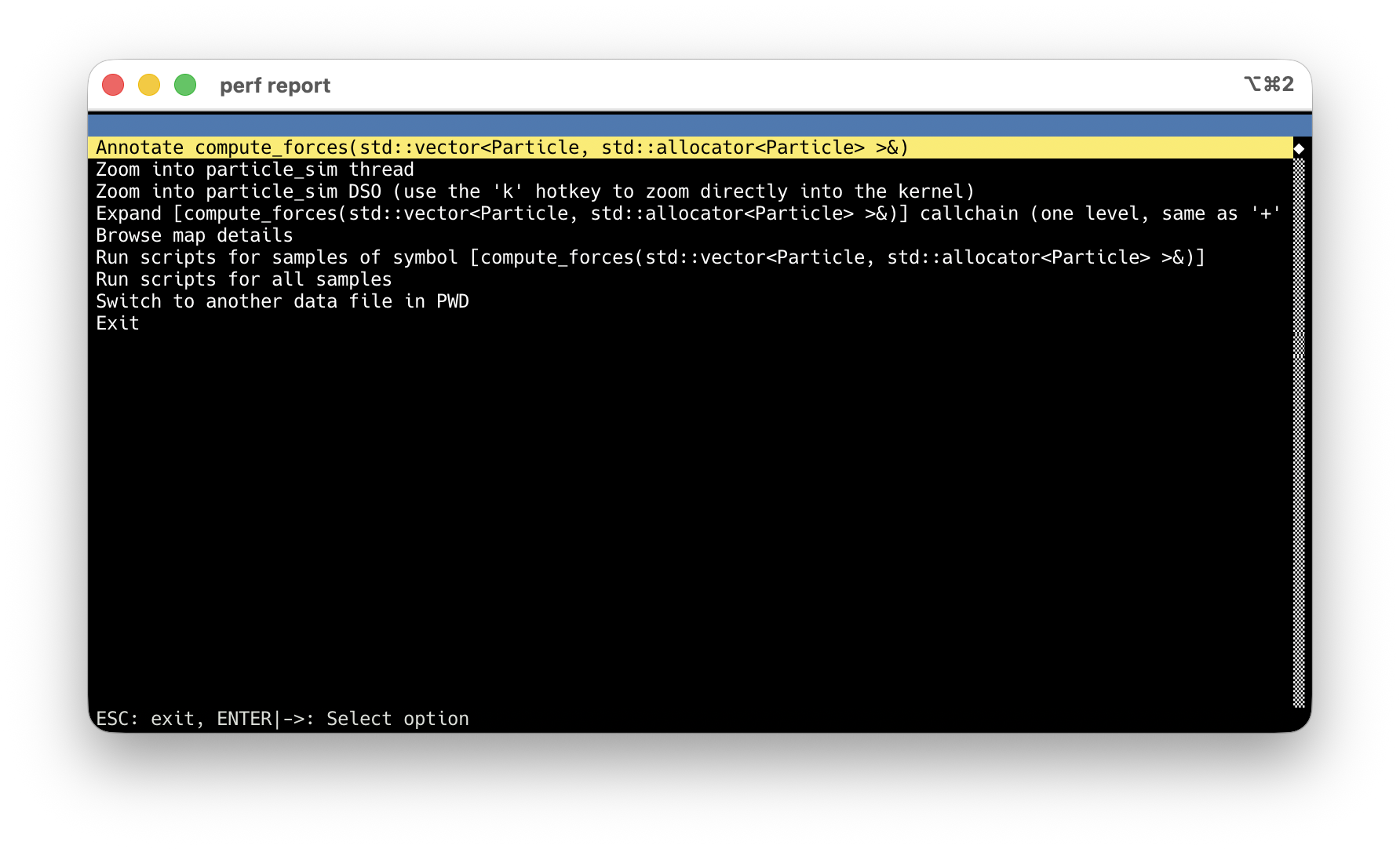
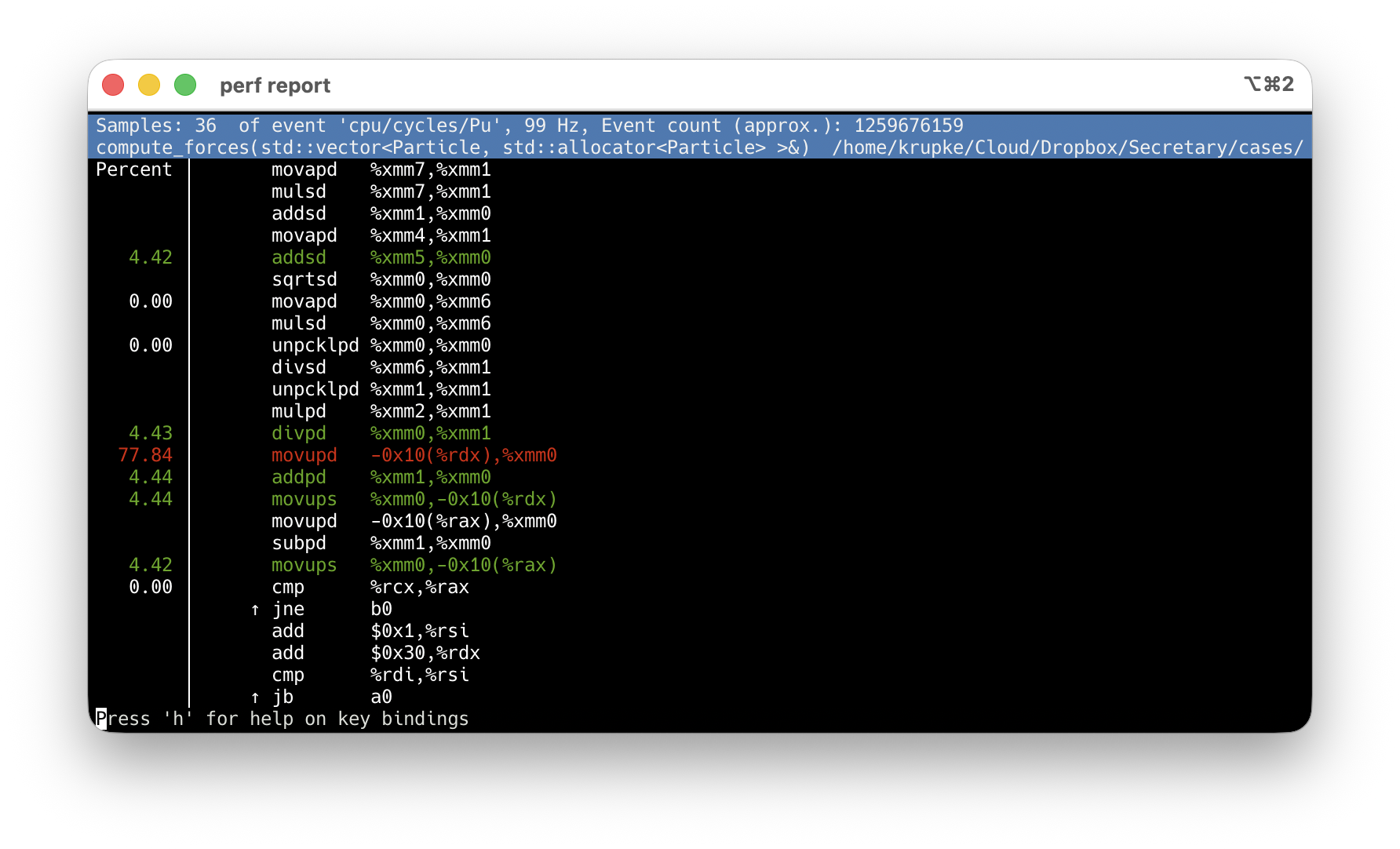
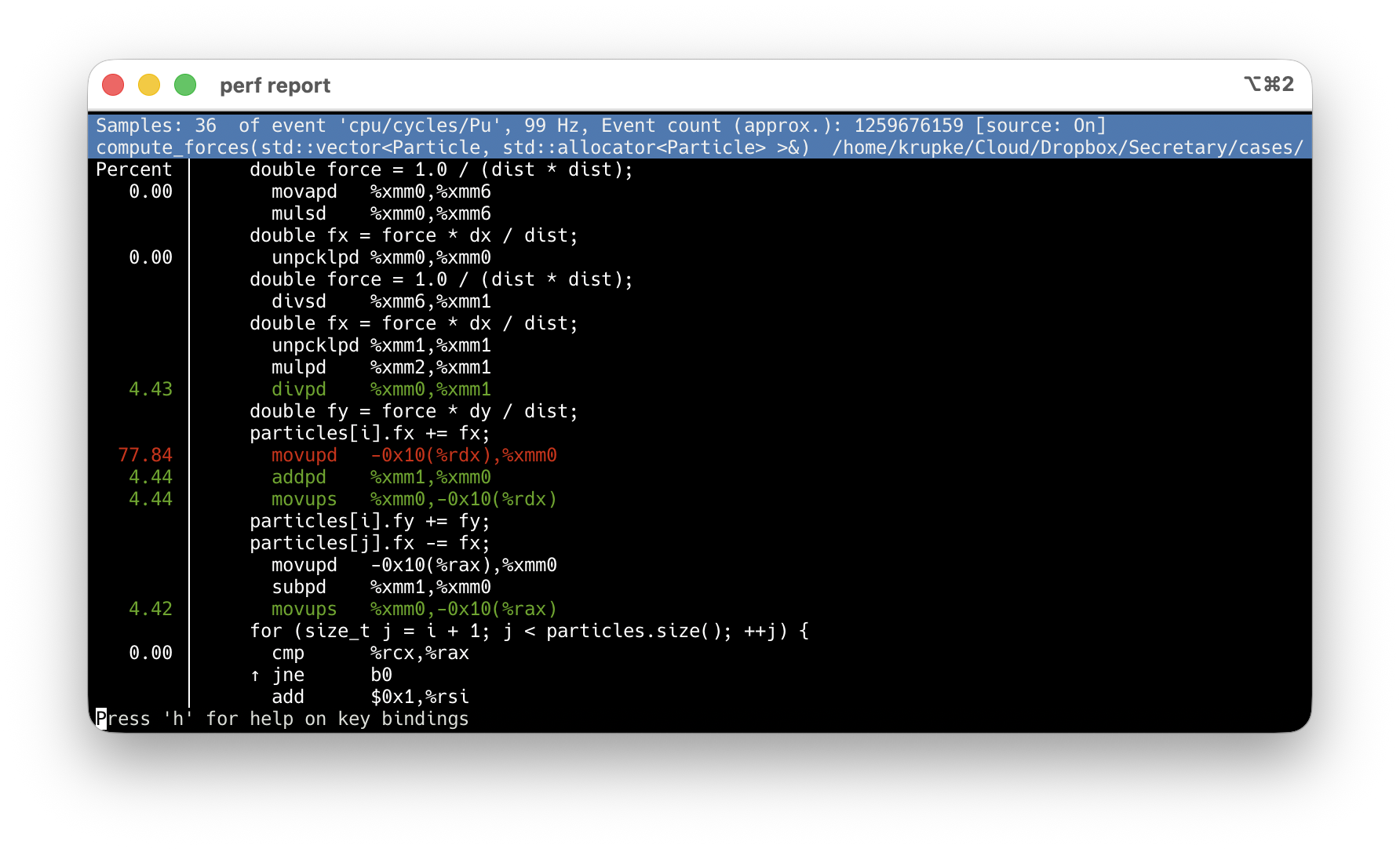
-O2 -g: debug info maps samples to source lines. Never use -O0.
compute_forces at 100% Children — every sample goes through it.
Arrow keys → highlight the hot symbol.
Enter → action menu → Annotate.
Default: asm-only, sample % per instruction.
Press s → source interleaved. Inlined code too.
Micro-benchmarking
Profiling finds where the bottleneck is. Micro-benchmarking measures whether your fix actually helped.
Python: timeit — Basic Usage (wall-clock time)
import timeit
n = 100_000
t_comp = timeit.timeit( # returns total seconds for `number` runs
lambda: [x**2 for x in range(n)], number=50
)
t_map = timeit.timeit(
lambda: list(map(lambda x: x**2, range(n))), number=50
)
print(f"comprehension: {t_comp:.3f}s map: {t_map:.3f}s ratio: {t_comp/t_map:.2f}x")C++: Google Benchmark — Our Example
We benchmark two graph representations with the same API:
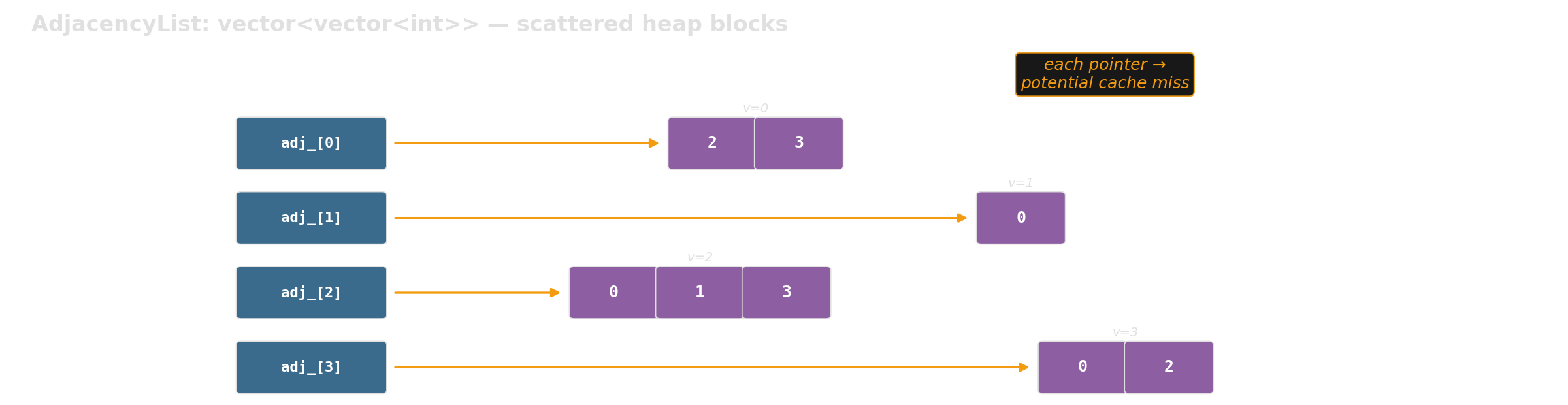
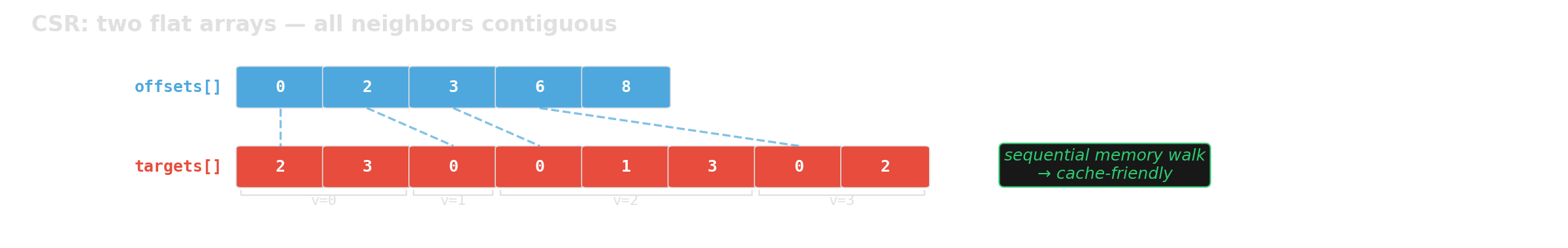
Which layout is faster for graph traversal at scale?
C++: Google Benchmark — Writing a Benchmark
static void BM_iterate_neighbors_csr(benchmark::State& state) {
int n = static_cast<int>(state.range(0)); // parameter from Range()
auto edges = random_edges(n, num_edges(n));
auto graph = make_csr(n, edges); // setup: NOT measured
for (auto _ : state) { // measured loop
long sum = 0;
for (int v = 0; v < graph.num_vertices(); v++) {
for (int u : graph.neighbors(v)) sum += u;
}
benchmark::DoNotOptimize(sum); // prevent dead-code elimination
}
state.SetItemsProcessed(state.iterations() * graph.num_edges() * 2);
}
BENCHMARK(BM_iterate_neighbors_csr)->RangeMultiplier(10)->Range(1000, 1000000);| Concept | What it does |
|---|---|
| Setup before loop | Excluded from timing |
for (auto _ : state) |
Framework controls iteration count |
DoNotOptimize(val) |
Prevents compiler from eliminating unused results |
SetItemsProcessed |
Enables throughput reporting (items/s) |
Range(lo, hi) |
Sweeps input sizes automatically |
Google Benchmark: Example Output
Benchmark Time CPU UserCounters
BM_iterate_neighbors_adjlist/1000 1,469 ns 1,465 ns 6.82G items/s
BM_iterate_neighbors_csr/1000 1,475 ns 1,472 ns 6.79G items/s
BM_iterate_neighbors_adjlist/1000000 14,541,591 ns 14,497 us 690M items/s
BM_iterate_neighbors_csr/1000000 8,369,209 ns 8,347 us 1.20G items/sAt N=1000: both fit in cache — no difference. At N=1M: CSR is 1.7x faster — memory layout matters.
Same semantics, different memory behavior — exactly the kind of result microbenchmarks should explain.
We have a plausible explanation — but perf stat can turn it into evidence.
Source: examples/google_benchmark/ — you will get this template.
Benchmark + perf stat: Why Is CSR Faster?
Filter to one case, wrap the binary with perf stat:
| Counter (per iteration) | adjlist | CSR | Ratio |
|---|---|---|---|
| time | 13.76 ms | 8.39 ms | 1.64× faster |
| instructions / cycle | 0.29 | 0.37 | 1.26× |
| L1-dcache miss rate | 8.1% | 5.6% | — |
| cache-misses (≈ DRAM) | 5.02 M | 1.80 M | 2.79× fewer |
Benchmark tells you how much. perf stat tells you why. Roughly 2.8× fewer DRAM-class misses — the pipeline is no longer starved.
Use the generic cache-references / cache-misses events — LLC-loads / LLC-load-misses report <not supported> on AMD Zen.
Setting Up Google Benchmark: Project Layout
my_project/
├── CMakeLists.txt # root: fetch deps, add subdirectories
├── src/
│ ├── CMakeLists.txt # defines the library target
│ └── graph/
│ ├── graph.h # your library headers
│ └── graph_gen.h
├── benchmarks/
│ ├── CMakeLists.txt # benchmark targets, link against library
│ └── bm_graph.cpp
└── tests/
├── CMakeLists.txt # test targets, link against library
└── test_graph.cppsrc/ defines a library target. Benchmarks and tests both link against it — same include paths, same compile settings, no relative-path #include hacks.
Setting Up: Root CMakeLists.txt
cmake_minimum_required(VERSION 3.14)
project(my_project LANGUAGES CXX)
include(FetchContent) # downloads libraries as part of cmake build
# Google Benchmark — fetched automatically, no manual install
FetchContent_Declare(googlebenchmark
GIT_REPOSITORY https://github.com/google/benchmark.git
GIT_TAG v1.9.4)
# Don't build benchmark's own tests (we don't need them)
set(BENCHMARK_ENABLE_TESTING OFF CACHE BOOL "" FORCE)
set(BENCHMARK_ENABLE_INSTALL OFF CACHE BOOL "" FORCE)
FetchContent_MakeAvailable(googlebenchmark)
add_subdirectory(src) # your library (INTERFACE target)
add_subdirectory(tests)
add_subdirectory(benchmarks)FetchContent downloads + builds Google Benchmark automatically. Just run cmake — no system-wide installation needed.
Full example with Google Test: examples/google_benchmark/CMakeLists.txt
Setting Up: benchmarks/CMakeLists.txt
add_executable(bm_my_algorithm bm_my_algorithm.cpp)
# Link against your library and Google Benchmark.
# my_lib provides include paths + compile settings (e.g., C++20).
# benchmark_main provides main() with built-in CLI flag parsing.
target_link_libraries(bm_my_algorithm my_lib benchmark::benchmark_main)Build and run:
Warning
Always use CMAKE_BUILD_TYPE=Release. Debug mode uses -O0 (no optimization) — your benchmarks will measure unoptimized code and the results are meaningless.
Full example: examples/google_benchmark/benchmarks/CMakeLists.txt — your existing src/ and tests/ stay untouched; benchmarks are additive.
Google Benchmark: Useful Flags
# Run only benchmarks matching a regex
./build/benchmarks/bm_graph --benchmark_filter=BFS
# JSON output for scripts/plotting
./build/benchmarks/bm_graph --benchmark_format=json \
--benchmark_out=results.json
# Multiple repetitions → mean, median, stddev
./build/benchmarks/bm_graph --benchmark_repetitions=5
# Combine: filter + repetitions + save to file
./build/benchmarks/bm_graph --benchmark_filter=iterate \
--benchmark_repetitions=5 \
--benchmark_format=json \
--benchmark_out=results.jsonAll flags are built into benchmark_main — no code changes needed.
Block 2: Achieve Performance
Now we know where the time goes. Here’s how to make it faster — in Python and in C++.
Python Performance Killers
Common patterns that are silently expensive. Each one is easy to fix — once you know it’s there.
x in list is secretly O(n)
Slow:
Building a temporary data structure just for faster lookups may feel redundant — but it is often the single most effective optimization.
Detour: std::set vs std::vector — not like Python
In Python, set always beats list for lookups (1000x). In C++, the picture is very different.
100k lookups against a container of size N:
| N |
unsorted vector O(n)
|
std::set O(log n)
|
sorted vector O(log n)
|
|---|---|---|---|
| 1K | 1.5 ms | 1.9 ms | 1.7 ms |
| 10K | 126 ms | 4.2 ms | 2.4 ms |
| 100K | 1,258 ms | 11.9 ms | 3.5 ms |
| 1M | 12,690 ms | 29.2 ms | 4.9 ms |
At N=10K, std::set beats the linear scan by only 30x — not 1000x like in Python. And the sorted vector beats std::set by up to 6x — same O(log n), fewer cache misses.
In C++, data layout often matters more than algorithmic complexity class. Prefer flat, contiguous containers unless insertions/deletions at scale make re-sorting too expensive.
String concatenation in loops
Strings are immutable in Python. Each += copies the entire string so far. That is \(O(n^2)\) in total length.
Use builtins instead of manual loops
Also applies to min(), max(), any(), all() — all run in C, not Python bytecode.
The general principle: push hot iterations into C-level code paths — the same idea as pybind11, just within the standard library.
Memoization with @lru_cache
Python’s @lru_cache makes this a one-line fix. Any pure function with repeated inputs is a candidate — DP subproblems, normalization, parsing.
Push numeric work to NumPy
Slow:
If Scalene shows high Python time on numeric loops, NumPy (or moving to C++) is the answer — not micro-optimizing the loop.
Data containers: cost of abstraction
Creating 100k objects with three float fields:
| Container | Creation | Create + access | Relative |
|---|---|---|---|
tuple |
1.0 ms | 11 ms | 1x |
class / dataclass |
15 ms | 23 ms | ~2x |
dataclass(slots=True) |
14 ms | 22 ms | ~2x |
NamedTuple |
18 ms | 27 ms | ~2.4x |
Pydantic BaseModel |
51 ms | 59 ms | ~5x |
Pydantic validates and coerces every field on every instantiation. That is invaluable at system boundaries — and ruinous in a hot loop.
Use Pydantic for parsing external input (API, config, files). Use dataclass(slots=True) or plain tuples for internal data that gets created millions of times.
Moving to Native: pybind11
The bottleneck is not “Python”. The bottleneck is the interpreted loop.
When pybind11 is worth it
- Scalene shows most time in Python execution
- the hotspot is a tight inner loop
- the Python-side implementation is fundamentally loop-heavy
- moving the whole loop is realistic
Our example: TSP nearest-neighbor + 2-opt local search. Both are tight O(n2) loops over distance computations — exactly the pattern where moving the entire loop to C++ pays off.
If the bottleneck is CPU-bound — especially superlinear algorithms — copying data into C++ and back is often worth it. The copy cost is linear; the savings are superlinear.
The measured difference
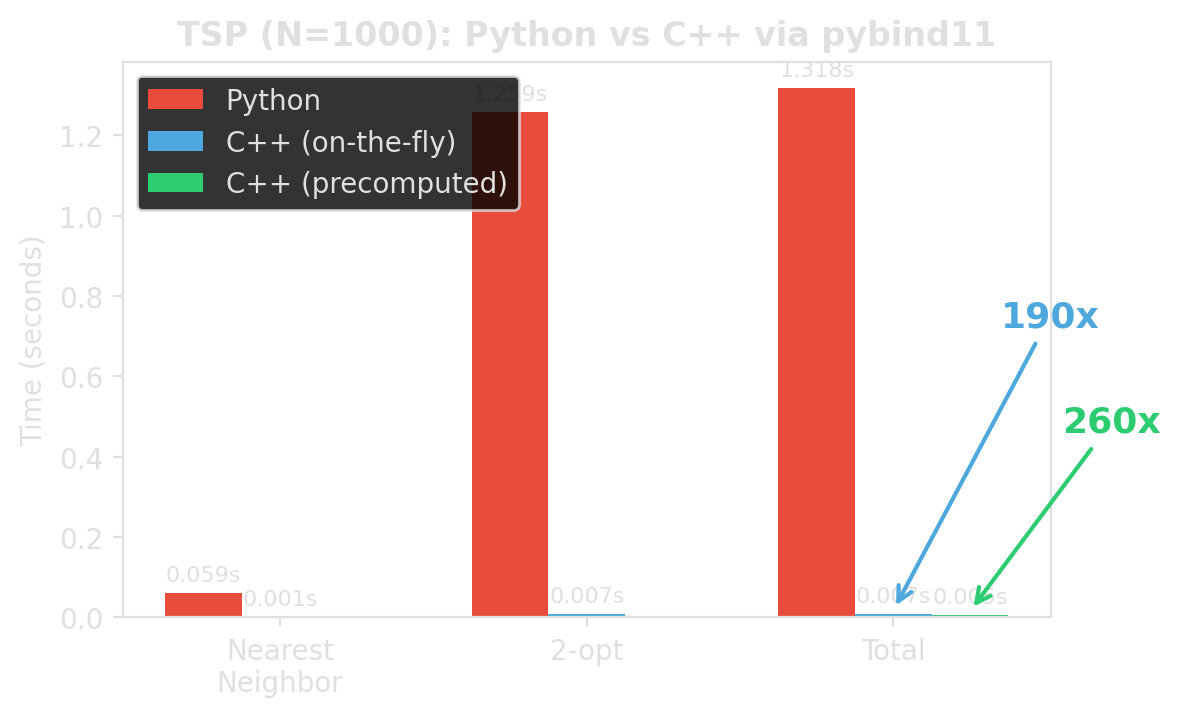
Same algorithm. Same result. Roughly 190x / 260x faster. Different place where the loop runs.
Source: examples/pybind11_template/benchmark_python_vs_cpp.py
The workflow
- Write the algorithm in pure C++ — no pybind11 dependency
- Write a thin binding file — converts Python types to C++ and back
- Configure the build —
pyproject.toml+CMakeLists.txt - Install with pip —
pip install -e .builds everything - Import from Python —
import fast_tsp; fast_tsp.nearest_neighbor(cities)
my_package/
├── pyproject.toml # build config (scikit-build-core + pybind11)
├── CMakeLists.txt # compile the C++ extension
├── src/my_package/
│ ├── algorithm.h / .cpp # pure C++ (no pybind11 dependency)
│ ├── _bindings.cpp # thin pybind11 wrapper
│ └── __init__.py # re-export C++ functions
└── tests/Step 1: Write the C++ algorithm
// tsp.h — pure C++, no pybind11 dependency
struct Point {
double x;
double y;
};
inline double dist(const Point& a, const Point& b) {
double dx = a.x - b.x;
double dy = a.y - b.y;
return std::sqrt(dx * dx + dy * dy);
}
std::vector<int> two_opt_improve(std::span<const Point> cities,
std::vector<int> tour) {
int n = static_cast<int>(tour.size());
bool improved = true;
while (improved) {
improved = false;
for (int i = 0; i < n - 1; i++) {
for (int j = i + 2; j < n; j++) {
double old_d = dist(cities[tour[i]], cities[tour[i+1]])
+ dist(cities[tour[j]], cities[tour[(j+1)%n]]);
double new_d = dist(cities[tour[i]], cities[tour[j]])
+ dist(cities[tour[i+1]], cities[tour[(j+1)%n]]);
if (new_d < old_d - 1e-10) {
std::reverse(tour.begin()+i+1, tour.begin()+j+1);
improved = true;
}
}
}
}
return tour;
}Full code: examples/pybind11_template/src/fast_tsp/tsp.h
Step 2: Write the binding file
#include <pybind11/pybind11.h>
#include <pybind11/stl.h> // auto-convert std::vector <-> Python list
#include <array>
#include "tsp.h"
namespace py = pybind11;
PYBIND11_MODULE(_native, m) {
m.doc() = "fast_tsp C++ extension — TSP routines via pybind11";
m.def("nearest_neighbor",
[](const std::vector<std::array<double, 2>>& cities) {
std::vector<Point> pts; pts.reserve(cities.size());
for (auto& c : cities) pts.push_back({c[0], c[1]});
py::gil_scoped_release release; // let other Python threads run
return nearest_neighbor(pts); // std::vector<int> → list
},
"Nearest neighbor heuristic",
py::arg("cities"));
m.def("two_opt_improve",
[](const std::vector<std::array<double, 2>>& cities,
std::vector<int> tour) {
std::vector<Point> pts; pts.reserve(cities.size());
for (auto& c : cities) pts.push_back({c[0], c[1]});
py::gil_scoped_release release;
return two_opt_improve(pts, std::move(tour));
},
py::arg("cities"), py::arg("tour"));
// Precomputed distance matrix for repeated access (e.g., multi-pass 2-opt)
py::class_<DistanceMatrix>(m, "DistanceMatrix")
.def(py::init([](const std::vector<std::array<double, 2>>& cities) {
std::vector<Point> pts; pts.reserve(cities.size());
for (auto& c : cities) pts.push_back({c[0], c[1]});
return DistanceMatrix(pts);
}),
py::arg("cities"))
.def_property_readonly("size", &DistanceMatrix::size)
.def("query", &DistanceMatrix::operator(),
py::arg("i"), py::arg("j"));
// Overloads taking DistanceMatrix instead of coordinates
m.def("two_opt_improve",
[](const DistanceMatrix& dm, std::vector<int> tour) {
py::gil_scoped_release release;
return two_opt_improve(dm, std::move(tour));
},
py::arg("distances"), py::arg("tour"));
m.def("nearest_neighbor",
[](const DistanceMatrix& dm) {
py::gil_scoped_release release;
return nearest_neighbor(dm);
},
py::arg("distances"));
}Source: examples/pybind11_template/src/fast_tsp/_bindings.cpp
Step 3: pyproject.toml + CMakeLists.txt
The build config has two pybind11-specific parts:
pyproject.toml — build system:
CMakeLists.txt — compile the extension:
cmake_minimum_required(VERSION 3.15...3.30)
project(${SKBUILD_PROJECT_NAME} LANGUAGES CXX)
set(PYBIND11_FINDPYTHON ON)
find_package(pybind11 3.0 CONFIG REQUIRED)
pybind11_add_module(_native
src/fast_tsp/tsp.cpp
src/fast_tsp/_bindings.cpp)
target_compile_features(_native PRIVATE cxx_std_20)
install(TARGETS _native DESTINATION fast_tsp)pip install -e . calls scikit-build-core, which calls CMake, which compiles the extension. The rest of pyproject.toml ([project] name, version, dependencies) is standard Python packaging.
Full files: examples/pybind11_template/pyproject.toml, CMakeLists.txt
Step 4: __init__.py and install
Use from Python:
Full template: examples/pybind11_template/
Distributing compiled extensions
pip install -e .works locally — but requires a C++ compiler + CMake on the target- Extensions are bound to a specific Python version and platform
- Wheels are pre-compiled —
pip installjust unpacks, no build tools needed - cibuildwheel + GitHub Actions automates building wheels for all platforms
For course projects, editable installs are fine. For packages you share, build wheels in CI so users never need a compiler.
C++ Performance Killers
Now the bottleneck is inside C++. Same principle: measure, then fix the right thing.
Accidental copies: auto r vs const auto& r
Slow:
One character (&) eliminates 100,000 heap allocations per iteration.
Missing reserve() for push_back
Slow:
Without reserve, the vector reallocates and copies ~24 times for 10M elements. Each reallocation touches an ever-larger memory region, thrashing the cache.
Erase-in-loop: O(n^2) hiding in plain sight
Slow:
Each erase() shifts all following elements. Repeated erases: \(\sum_{k=1}^{n} O(n-k) = O(n^2)\).
Memory layout: AoS vs SoA
AoS (Array of Structs):
AoS reads 32 bytes per element but uses only 4. 7/8 of cache bandwidth is wasted.
Why AoS wastes cache bandwidth
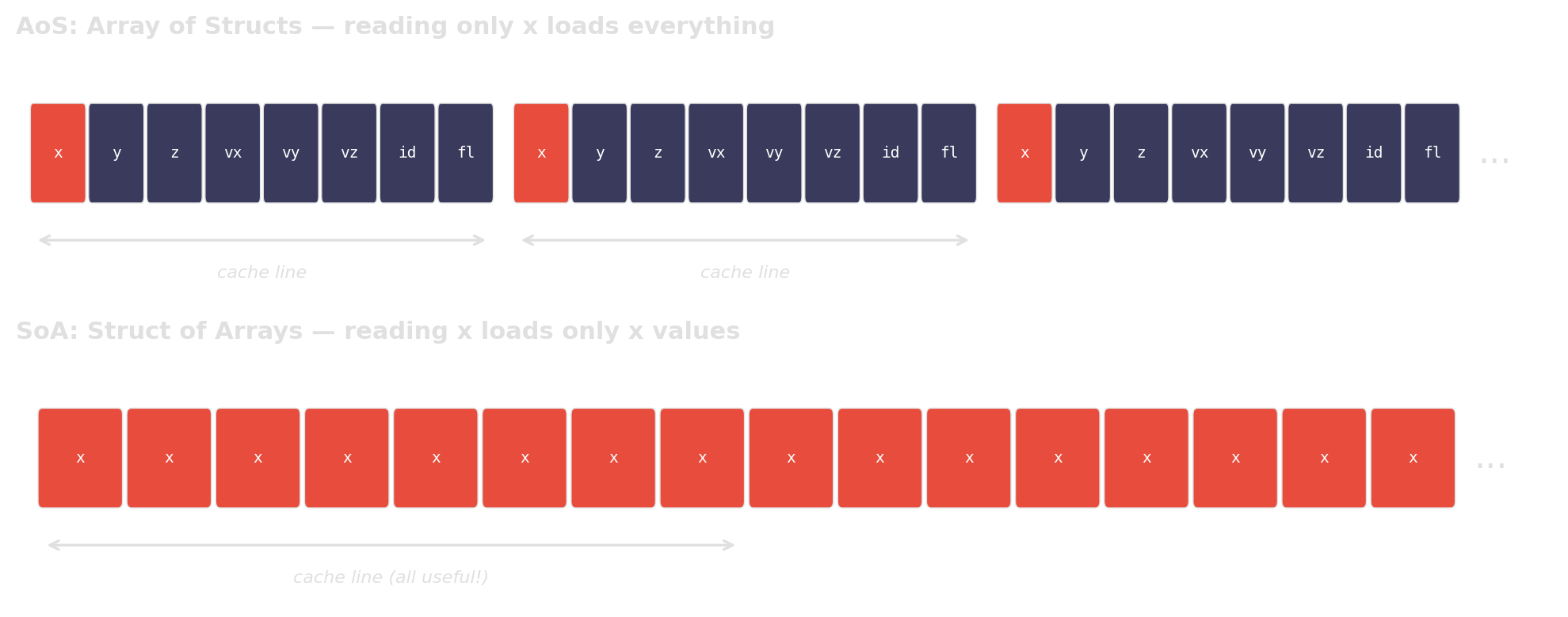
When you only access one field, AoS forces the CPU to load the entire struct per element. SoA packs the relevant data contiguously, so every cache line is fully utilized.
Pointer chasing: std::list vs std::vector
Slow:
Sequentially allocated nodes happen to sit nearby in memory — best case. In real programs, nodes are interleaved with other allocations — worst case.
Every node->next is a potential cache miss. A flat array lets the prefetcher do its job.
Accidental \(O(n^2)\): dedup with nested loops
Slow:
Algorithmic thinking is not just for Python. The same complexity traps appear in C++ — the fix is often a standard-library one-liner.
Wrap-up
The discipline matters more than the specific tools.
Same discipline, every time
- Find the dominant cost — logging, then profilers
- Classify the bottleneck — algorithmic? interpreter overhead? memory layout?
- Change one thing — the thing the evidence points to
- Verify on the same workload — timeit, Google Benchmark, or just timestamps
pybind11 is not “make it fast”. Google Benchmark is not “time it nicely”. Both are part of a measurement-driven workflow.
Concepts, not tools
| Tool | Concept | Alternatives |
|---|---|---|
logging |
Structured observability | loguru, spdlog (C++) |
| Scalene | Sampling profiler (Python) | py-spy, VTune |
perf |
Sampling profiler (C++) | VTune, Instruments (macOS) |
| cProfile | Tracing profiler | yappi, gprof |
| pybind11 | Native bindings for Python | nanobind, cython, PyO3 |
timeit |
Microbenchmarking (Python) | pytest-benchmark |
| Google Benchmark | Microbenchmarking (C++) | Catch2, criterion (Rust) |
Further reading
- Performance Analysis and Tuning on Modern CPUs — Denis Bakhvalov (CC0, free). Companion exercises: perf-ninja.
- What Every Programmer Should Know About Memory — Ulrich Drepper (2007). The classic on memory hierarchy and caches.
- Data Structures in Practice — Danny Jiang (CC BY 4.0). Hardware-aware data structures, backed by
perfbenchmarks.
- Catching Cache: A Detective Story — Michelle Nguyen (Bloomberg, ~42 min). Why
vectorbeatsunordered_mapby 10×. - Matrix Multiplication Optimization — Alexia Salah (Codeplay, ~46 min). 150× speedup: loop order → tiling → SIMD → cache-aware design.
- Want fast C++? Know your hardware! — Timur Doumler (CppCon 2016, ~60 min). Caches, branch prediction, false sharing, denormals.
Links
| C++ Core Guidelines | Parameter passing, copy semantics, best practices |
| Google Benchmark | Microbenchmarking framework for C++ |
| pybind11 Documentation | C++/Python bindings — reference & tutorial |
| Course Material Repository | Slides, examples, and exercises |
References
Algorithm Engineering SS 2026 — T01 Performance Engineering